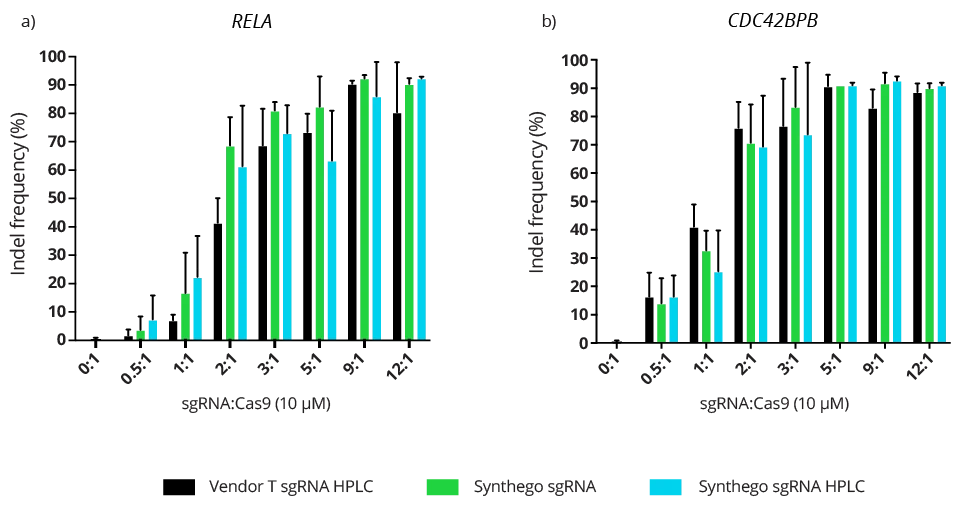
Not all chemically synthesized sgRNA are manufactured equally. During synthesis, some oligomers fail to extend during each cycle resulting in undesired truncated molecules mixed in with your desired sgRNA. Additionally, several types of chemical contaminants can be introduced through the manufacturing process of sgRNA. The presence of both these impurities can increase off-target binding, reduce on-target binding, or both. Several vendors offer high-performance liquid chromatography (HPLC) as a purification step to eliminate the unwanted truncated oligomers (n-3 to n-5) and contaminants. However, each vendor might have different synthesis and quality standards to reach the purity levels desired for your sgRNA.
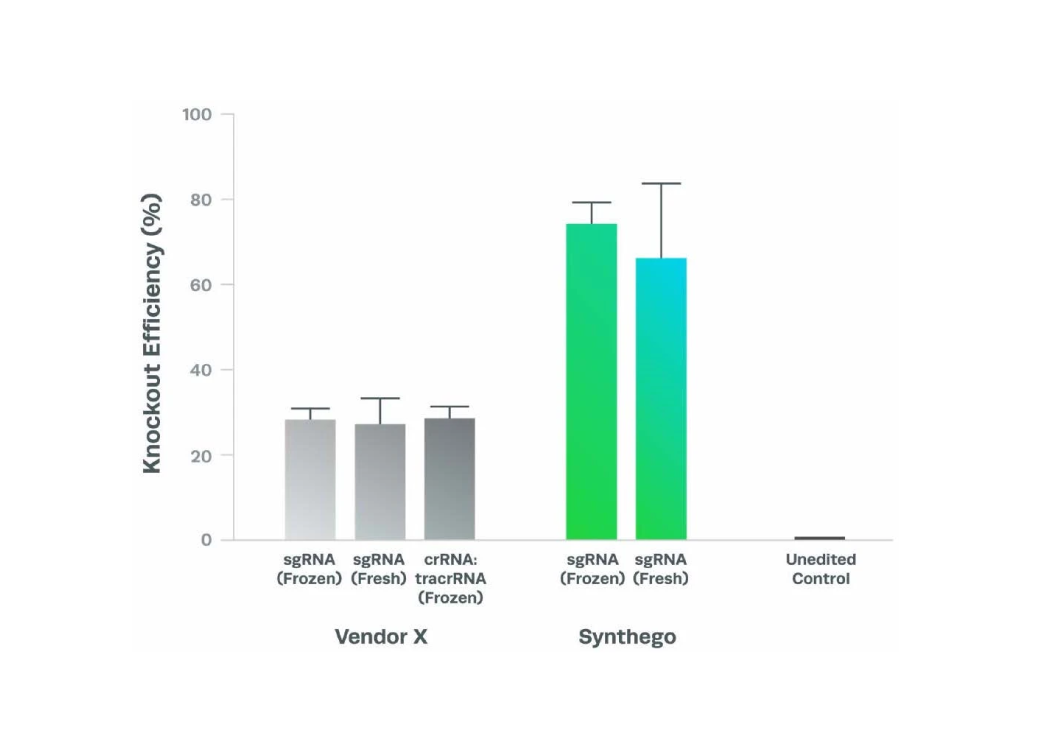
Synthego’s best-in-class sgRNA can achieve exceptional editing efficiencies through standard manufacturing processes (non-HPLC) compared to HPLC purified sgRNA from other vendors. If the purity was substantially better, you would expect the HPLC purified sgRNA to perform better in all cases. However, that is not always the case in our study.
Depending on your application, particularly sensitive applications like therapeutic development require sgRNA that achieves high performance in early development and highly pure sgRNA as development progresses towards the clinic.