The following modifications are included on our hfCas12Max modified gRNA:
2'-O-Methyl analog at the first 3 bases. The last 4 bases have 3 modifications ending with a nonmodified base. With 3' phosphorothioate bonds between 3 first and last 4 bases.
Gene Editing With High Precision
hfCas12Max mRNA encodes a high-fidelity Cas12 nuclease engineered for precise genome editing with reduced off-target activity. Transient mRNA delivery enables controlled nuclease expression while supporting efficient editing when paired with compatible CRISPR guide RNAs.
hfCas12Max mRNA encodes a high-fidelity Cas12 nuclease for precise genome editing with expanded target access. Designed for use in mRNA-based delivery workflows, it supports transient nuclease expression, broad PAM recognition, and flexible application across diverse cell types.
High-Fidelity Editing:
Engineered Cas12 variant delivers strong on-target performance with minimized off-target cleavage.
Transient Expression
Supports time-limited nuclease activity, enabling precise control over genome editing windows.
Broad Targeting Capability:
PAM recognition expands access to diverse genomnic loci across multiple applications.
Flexible Delivery Compatibility:
Designed for use across multiple delivery approaches.
| Product Number | M20HFCAS12MAX |
| Concentration | 1 µg/µL |
| Delivery Format | Tubes |
| Intended Use | This product is intended for research use only |
| Source | in vitro transcription |
| Shipping Conditions | Dry Ice |
| Storage Temperature | -80 C |
| Buffer Composition | 1 mM Sodium Citrate, pH 6.4 |
| Function | Enables high-fidelity gene editing with an expanded targeting range through direct delivery of a ribonucleoprotein (RNP) complex |
| For Use In | CRISPR gene editing experiments utilizing the hfCas12Max nuclease mRNA |
| Specification | hfCas12Max Nuclease |
|---|---|
| Size | 3672 nt |
| PAM Sequence (N = any nucleotide) | 5'-TN-3' or 5'-TTN-3' |
| DNA Cleavage | Staggered-cut: cleavage on the target strand occurs 24 nt downstream from the PAM, while the non-targeted strand is cut 14-16 nt downstream |
| Endonuclease Domains | RuvC |
| gRNA Length | 44 - 50 nt (crRNA only, no tracrRNA) |
| Target Sequence Length | 20 nt |
| Enzyme Class | Type V CRISPR-Cas system of Cas12 |
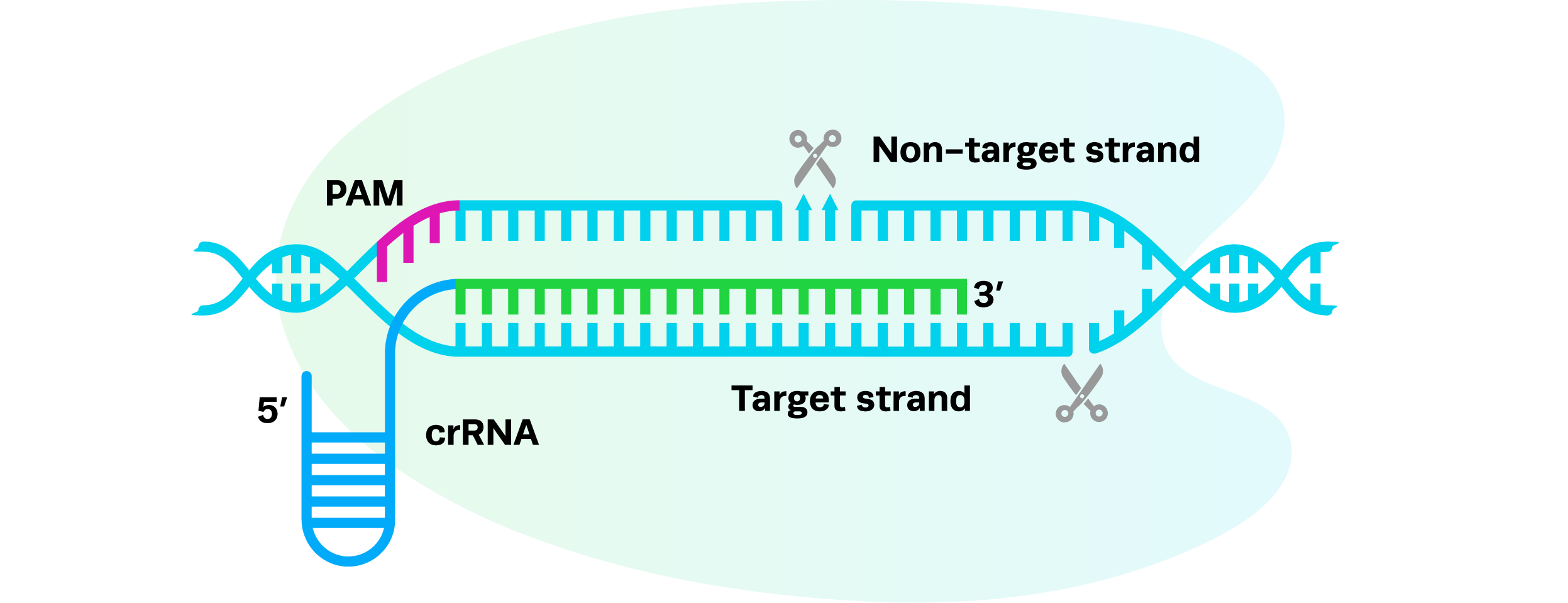
hfCas12Max utilizes a single crRNA, eliminating the need for a tracrRNA. The guide RNA is compact (44–50 nt), including a 17–23 nt target sequence complementary to the genomic locus of interest. When hfCas12Max is expressed intracellularly from mRNA, the nuclease associates with the crRNA to enable targeted DNA cleavage, generating staggered cuts downstream of the PAM site.
For guide design, target sequences should be selected adjacent to a 5'-TTN-3' PAM. Designed sequences can be generated using standard CRISPR guide design tools and synthesized with optional chemical modifications to support stability and editing performance in cellular workflows.
Try hfCas12Max nuclease and gRNA in your workflow with our validated* gRNAs below.
| Gene Name | hfCas12Max Target Sequence |
|---|---|
| B2M | TATCTCTTGTACTACACTGA |
| TRAC | GAGTCTCTCAGCTGGTACAC |
The following modifications are included on our hfCas12Max modified gRNA:
2'-O-Methyl analog at the first 3 bases. The last 4 bases have 3 modifications ending with a nonmodified base. With 3' phosphorothioate bonds between 3 first and last 4 bases.
Legal Terms: hfCas12Max Nuclease purchased from Synthego is only intended and licensed to be used solely with hfCas12Max guide RNAs purchased from Synthego, and hfCas12Max guide RNAs purchased from Synthego are only intended and licensed to be used solely with hfCas12Max Nuclease purchased from Synthego. For more details, visit our patent page.