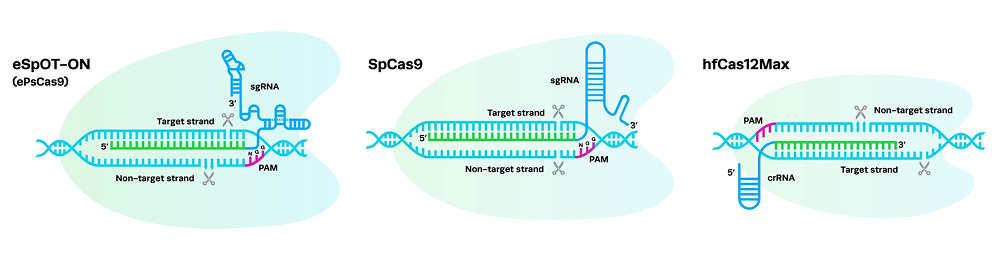
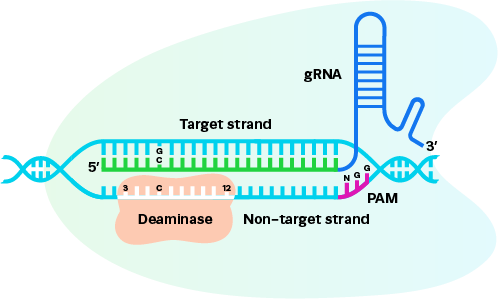
Consistent manufacturing and documentation strategies that support seamless progression from early discovery to regulated workflows.
→
Learn more
Webcast featuring eSpOT-ON
Explore More
eSpOT-ON Nuclease Protein Available Now
Explore More
Order eSpOT-ON Nuclease mRNA Now