To this date, Cas12 and Cas13 systems are the primary drivers of signal generation in CRISPR-based assays due to their collateral cleavage activity. Cas12 is most commonly used for DNA detection following RPA amplification, while Cas13 enables RNA detection in both direct and amplification-coupled workflows. For DNA-targeted detection workflows, engineered Cas12 nucleases are often explored to improve signal-to-noise performance. Synthego’s hfCas12Max, a high-fidelity Cas12i-family nuclease, is designed to reduce non-specific collateral activity while retaining target-triggered activation and can be evaluated in CRISPR-based detection workflows.
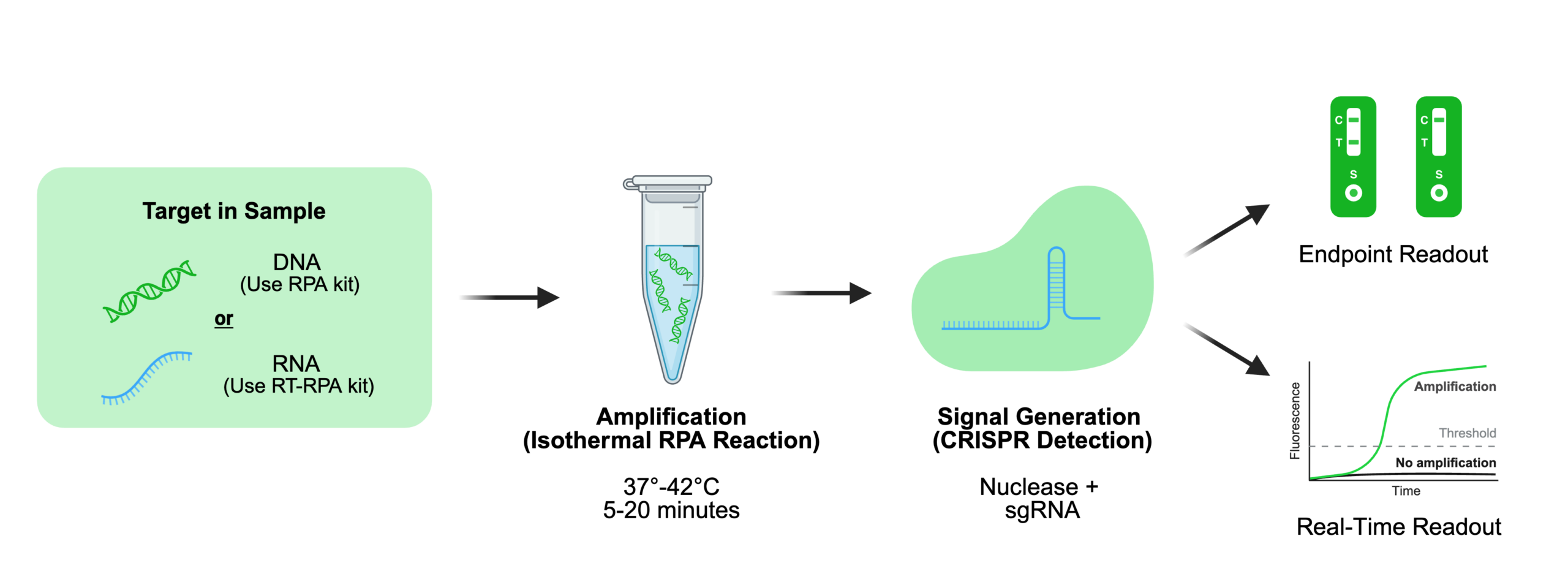
Cas 12 and Cas13 systems convert target recognition into measurable outputs through fluorescent reporter cleavage. Fluorescent reporters support real-time or quantitative detection, while lateral flow formats enable rapid, instrument-free readouts. In lateral flow systems, signal interpretation depends on reporter cleavage state, where intact and cleaved reporters produce distinct visual outputs. The choice between endpoint and real-time detection ultimately depends on assay requirements such as sensitivity, quantitation, and available instrumentation, with each format offering trade-offs in speed, complexity, and data resolution. For a deeper comparison, see our endpoint versus real-time RPA guide.
In contrast, Cas9 lacks collateral activity and is typically used in engineered detection systems where signal generation is externally coupled. While this makes Synthego enzymes like SpCas9 and high-fidelity variants such as eSpOT-ON exceptionally precise and reliable for applications like gene editing—where controlled, site-specific cleavage is essential—it limits their utility in diagnostics. Without intrinsic signal amplification through collateral cleavage, Cas9-based assays often require additional components or steps, increasing complexity and potentially reducing speed and sensitivity compared to systems built around enzymes like Cas12 or Cas13.